Research
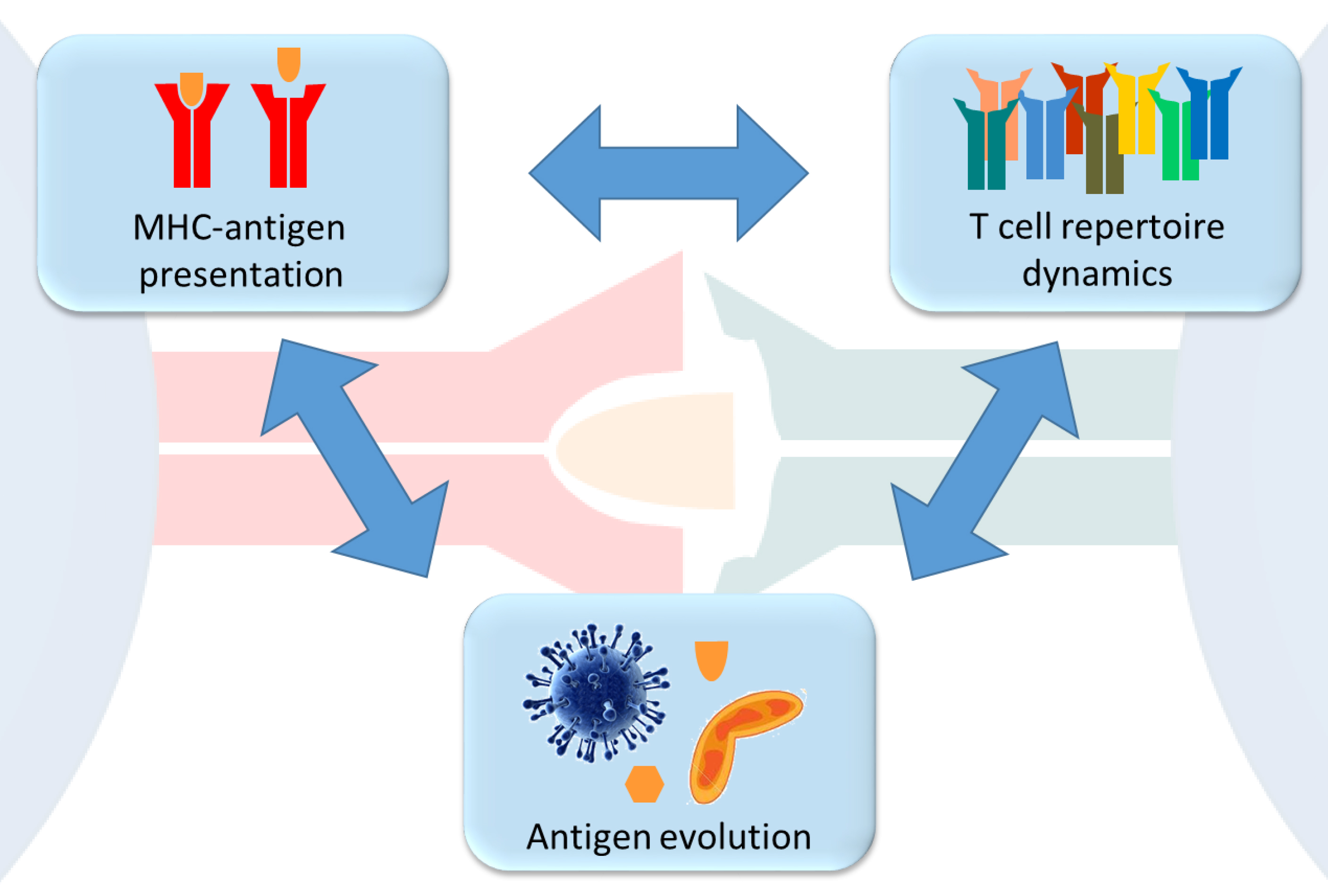
Our research investigates the evolutionary origin and consequences of functional genetic diversity in the context of health and disease. A particular focus of the group lies on the dynamics and trade-offs that govern successful antigen presentation and recognition in the adaptive immune system of vertebrates. We are exploring the complex interactions between the functional variability of antigen-presenting MHC molecules, the dynamics of the interacting T cell repertoire, and the antigen evolution in pathogens in order to improve our understanding of immune-mediated diseases in particular and the evolution of the adaptive immune system in general.
