Research
A better understanding of molecular mechanisms that safeguard the specific characteristics and the health of cells is essential for basic research as well as for biomedical applications.
Cell fate specification processes during development have been elucidated in great detail. Yet, there is a lack of knowledge regarding molecular safeguarding mechanisms that protect cell identities of differentiated cells and maintain their function and health. Knowledge about cellular safeguarding mechanisms will improve approaches to generate fresh cells for prospective regenerative medicine applications.
In this context, important aspects of safeguarding mechanisms are how they counteract cellular reprogramming and how Aging is affected when safeguarding is weakened or lost. Overall, a better understanding of the complex molecular networks and pathways that contribute to maintaining and safeguarding cell identities, function, and health will support using reprogramming to generate stem cells and specific tissues for future regenerative therapies.
Research Subjects
C. elegans to elucidate molecular mechanisms of cellular safeguarding
We use the nematode Caenorhabditis elegans as a genetic model organism to investigate the molecular basics of cellular safeguarding. Sydney Brenner established around 1963 the roundworm C. elegans as a genetic model organism for biological research. Since then, C. elegans has proven to be a highly efficient model organism for a broad spectrum of questions in Life Sciences. C. elegans was the first metazoan organism with a fully sequenced genome. Since approximately 60% of all C. elegans genes have human homologs, many regulatory processes are conserved in higher organisms and humans. This is reflected by the fact that several C. elegans researchers pioneered discoveries including developmental genes, RNA interference (RNAi), programmed cell death (Apoptosis), aging regulation, the first micro RNAs lin-4 / let-7 - just to name but a few - leading to several Nobel Prize awards. Such achievements emphasize the high potential of C. elegans as a model organism to help understanding fundamental questions in basic as well as biomedical research. We apply a varies of molecular and cell biological techniques, including transcriptome analysis (RNA-Seq), single-molecule fluorescent in situ hybridization (smFISH), chromatin-Immuno-precipitations with sequencing (ChIP-Seq for transcription factors, chromatin factors, and Histone modifications). In addition, we use Mass Spectrometry to identify protein-protein interactions (CoIP-MS) and measure changes in metabolism. Furthermore, we assess chromatin signatures and accessibility using ATAC-Seq and validate biochemical interactions in vivo by assessing genetic interactions. Applying CRISPR/Cas9-mediated gene editing allows us to modulate gene expression and perform tissue-specific protein biochemistry. Overall, our goal is to better understand how epigenetic mechanisms, metabolism, physiological changes, and infections influence cellular safeguarding in the context of reprogramming and Aging.
Epigenetics in the context of cellular safeguarding, reprogramming, and Aging
One promising source for fresh cells to replenish and repair diseased organs is converting abundantly available cells to the required cell type. Such conversion can be achieved by reprogramming upon overexpression of specific transcription factors. Yet, transcription factors are restricted in their efficiency to induce reprogramming due to cell fate maintenance and safeguarding mechanisms. The resulting limitations in reprogramming efficiency are critical in particular for in vivo applications. To overcome these limitations, it is vital to better understand the mechanisms that safeguard cell identities and thereby impose barriers for reprogramming. Repressive epigenetic signatures based on histone modifications and chromatin conformation can impose barriers for reprogramming by preventing transcription factors from activating their target genes. Histone-modifying and chromatin-binding proteins are often influenced by a variety of different factors such as chaperones, kinases, RNA-binding proteins, and metabolites. Therefore, we elucidate molecular pathways that integrate different metabolic and cellular states to elicit changes in epigenetic gene expression regulation.
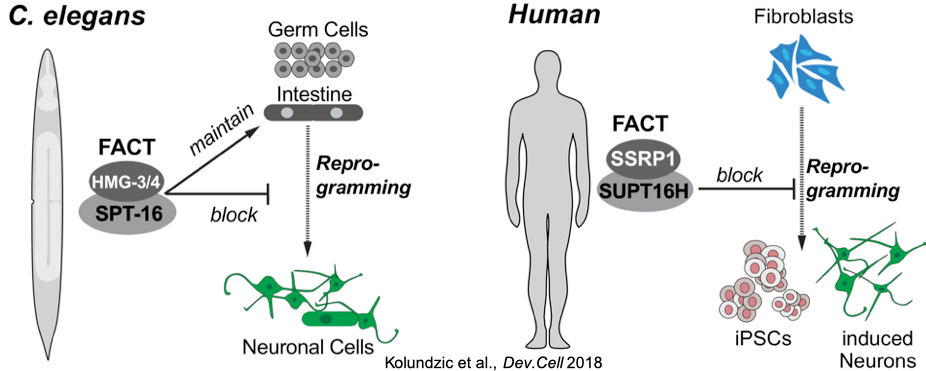
We established C. elegans as a model organism to study reprogramming in vivo, which allows genetic approaches to identify cell fate-safeguarding factors (Tursun et al., 2011 Science; Patel & Tursun et al.,2012 Cell Rep.; Seelk et al., 2016 eLife; Kolundzic et al., 2018 Dev. Cell; Hajduskova et al., 2019 Genetics; Kazmierczak et al., 2021 NAR). Initially, we discovered the chromatin regulator LIN-53 (CAF-1p48/RBBP4 in humans) as a barrier for direct reprogramming of germ cells to neurons. Subsequently, other research groups recapitulated our findings with mammalian cells, demonstrating the conservation of safeguarding mechanisms in evolution. Recently, we also showed that the histone chaperone complex FACT blocks reprogramming in C. elegans as well as in human fibroblasts, thus providing another instance of reprogramming barrier conservation across species. Notably, some epigenetic factors, such as LIN-53, that act as cell fate safeguards are also implicated in aging regulation (Müthel et al., 2019 Aging Cell). We are pursuing this line of research by assessing to which degree loss of cellular safeguarding factors also affect Aging.
Epigenetics and metabolism are linked in the context of cellular safeguarding.
We make use of C. elegans as a powerful model organism for genetic screens to identify inhibitors of transcription factor-induced cell fate reprogramming. By performing reverse genetics (RNAi) screens, we identified epigenetic factors, proteostasis regulators, cytoskeleton components, and mitochondrial proteins, whose depletions create permissiveness for cellular reprogramming. Regarding the latter, we found that metabolism perturbations elicit changes in chromatin modifications and affect gene expression. As metabolome analysis (GC-MS-based) together with genetics point to the implication of key metabolites and chromatin regulators, we are studying the phenomenon of linked metabolism and epigenetic regulation in the context of cellular safeguarding. This concept is also illustrated by our recent study, where we applied metabolomics to reveal that epigenetic regulation of Trehalose levels by the histone chaperone LIN-53 is critical for lifespan regulation (Müthel et al., 2019 Aging Cell). Overall, our goal is to decipher the molecular pathways, which link metabolism and epigenetics to maintain cell fates and counteract the conversion of cell identities. Processes to compensate perturbed metabolism may ensure cellular and organismal survival but appear to weaken cell fate stability. This phenomenon relates to metabolic disorders but also to infections and tumor biology in the context of cancer cell survival. Generally, revealing how changes in metabolism or infections affect cellular safeguarding is instrumental for regenerative medicine as prospective approaches may include reprogramming of cells directly in patients suffering from different diseases.
Shutdown of cell identities
An essential aspect of reprogramming is that faithful conversion to another identity requires robust repression and erasure of the initial cell identity. We termed this process starting cell fate shutdown. It is plausible that safeguarding mechanisms counteract cell fate shutdown processes, thereby decreasing the efficiency of producing faithfully reprogrammed cells. In-depth transcriptome analysis of reprogrammed cells revealed that varying degrees of gene activities belonging to the starting cell fate often persist, which can limit the capacity of iPSCs to form terminally differentiated tissues. Additionally, during direct reprogramming of fibroblasts to neurons or to hepatocytes, a significant fraction of converted cells still expressed fibroblast genes. Such persisting gene expression may be detrimental by leading to tumor formation. Because reprogramming studies primarily focus on the induction of the new target cell, fate shutdown mechanisms remain elusive. Their understanding is crucial to increase the functionality and safety of reprogrammed cells for applications in regenerative medicine.
We recently discovered a novel reprogramming phenomenon in C. elegans to study cell fate shutdown in vivo. It allows investigating whether direct reprogramming of the same cell type to either neurons or intestinal cells follow common or separate trajectories with regard to shutting down the original fate. Besides dissecting molecular mechanisms in vivo, C. elegans provides the possibility to reveal cell-non-autonomous effects, which often remain undetected when studying cultured cells without the physiological context of different tissues and organs. To understand shutdown mechanisms better, we study to which degree transcription factors used for reprogramming are directly implicated and whether processes such as metabolism and chromatin regulation are involved.
