Plant Immune Biology / Research
Statement of Objectives
At the department of Plant Immune Biology, we adopt a multidisciplinary approach to understand how plants endure and overcome stress caused by infections. We aim to contribute in the development of sustainable agricultural practices, by understanding cellular processes at a molecular level, to develop strategies to improve crop resistance and thus contributing to "Meeting global challenges".
Our mission encompasses empowering students and early-career researchers through curiosity and inquiry-based learning experiences in plant biology. We aim to cultivate their expertise in molecular plant biology, biochemistry, and genetics, enabling them to explore the intricate workings of plants.
Research overview
At the Department of Plant Immune Biology, we investigate how plants are able to cope with biotic stresses. Plants are able to sense infections and respond to them rapidly in order to efficiently defend themselves. Responses to challenges occur at multiple levels, and coordinated proteolysis is paramount to allow plants to reshape the proteome, as well as to maintain protein homeostasis (proteostasis) during infection. We take a multidisciplinary approach to understand how the signalling molecule ubiquitin shapes cellular responses to stress.
We believe that understanding the basic mechanisms by which plants are able to survive challenging environments, will provide the basis for the development of solutions to enable sustainable agriculture in the face of climate change.
Our research areas
Mechanisms of ubiquitination
Despite the ubiquity of ubiquitination, it remains largely unknown how E3 ligases associate with various kinases to regulate cellular responses or which specific E2 enzymes determine the "ubiquitin code" that dictates protein fate, a gap our research addresses through newly developed tools for analyzing E2-E3 biochemical activities.
Even though ubiquitination participates in most, if not all cellular processes, many questions in the field remain unanswered. These include questions regarding the basis of the ubiquitination process. The final step is carried out by the E3 ubiquitin ligase by guiding the modification of a substrate, bringing both the E2-ubiquitin conjugate and the substrate together. One of the outstanding questions concerns the regulation of the E3 and E2 activity. Many U-box type E3 ligase such as PUB22 (Trenner et al. 2022), form signalling modules with different types of kinases, including receptor and mitogen-activated protein (MAP) kinases to regulate a plethora of cellular responses ranging from immunity to development (Trujillo 2021).

UHH/Trujillo
It remains largely unknown which type of ubiquitin chains are built by specific E2 conjugating-enzymes that largely determined chain linkage (e.g. Lys63 or Lys48-linked chains). The different ubiquitin polymers constitute the ubiquitin code, and determines different fates of the modified protein, such as degradation (Lys48) or changes in subcellular localization (Lys63). To uncover the biochemical activities of E2-E3 pairs we developed a set of tools that enables us to analyse the activities (Turek et al. 2018; Kowarschik 2018; Saeed et al. 2023).
Regulatory functions of ubiquitin
Focusing on the immune response, our research aims to uncover how ubiquitin signalling maintains cellular homeostasis by rerouting vesicle trafficking to regulate immune signalling.
Ubiquitin signalling plays a key role in the regulation of cellular responses. Our aim is to uncover cellular processes and the components that are targeted by ubiquitination, with a focus on the immune response (Trujillo 2021). Ubiquitin and ubiquitin-like protein modifiers play key roles in controlling signalling amplitude and intensity, as well as in buffering proteome imbalances caused by pathogen attack. An intertwined and complex signalling circuitry, regulates cellular dynamics and protein degradation to maintain homeostasis.

UHH/Trujillo
Our studies have revealed that PUB ligases interact with components of the vesicle trafficking. These include subunits of the exocyst, an essential complex that mediates the tethering of post Golgi vesicles to the plasma membrane required for the secretion of immune receptors (Brillada et al. 2021; Stegmann et al. 2012; Stegmann. et al 2013). During the immune response, subunits of the exocyst are phosphorylated and ubiquitinated inhibiting interaction with the plasma membrane, while inducing interaction with the autophagy machinery to reroute the exocyst to the vacuole to be degraded. These mechanisms control receptor delivery acting as molecular brakes to dampen of cellular responses and maintain homeostasis (Trujillo et al. 2008).
Ubiquitin SynBio
One of our ultimate aims is to engineer systems and new small molecules that mediate Targeted Protein Degradation (TPD). For this we are developing cost-effective and scalable platforms to analyse degradation processes.
The study of ubiquitinated proteins faces many challenges such as the low stoichiometry of ubiquitin, rapid cleavage of ubiquitin moieties and the degradation of modified proteins. By reconstituting the ubiquitination cascade in an orthogonal system such as E. coli, we are able to circumvents some of these limitations.
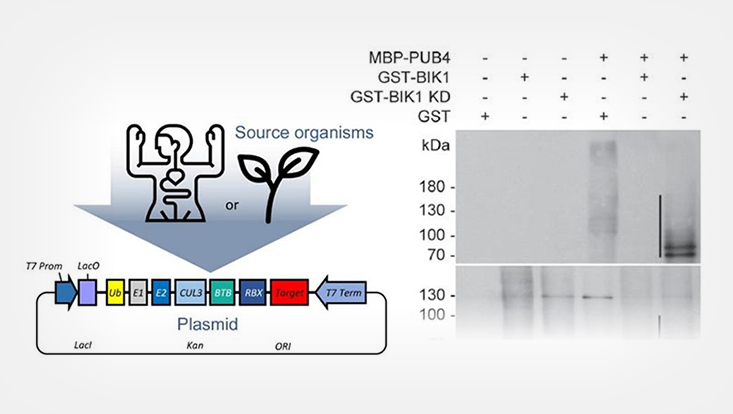
UHH/Trujillo
For this purpose, we developed a synthetic biology toolbox that is based on golden gate cloning (Kowarschik et al. 2018). Expression operons containing all components of the ubiquitination cascade, enable the easy analysis E3 activity or substrate ubiquitination, which can be expensive and time-consuming (Winkler et al. 2017; Turek et al. 2018; Kowarschik et al. 2018; Yu et al. 2023). Furthermore, UbiGate is also amenable for easy up-scaling, rendering ubiquitinated products accessible to further down-stream analysis such as mass-spectrometry or NMR.
